Overview
David King is a computational biologist whose work centers on the genetic underpinnings of complex biological traits. He builds the pipelines and develops the analyses that advance the biological question at hand. Learn more about his background at About Me.
Featured projects
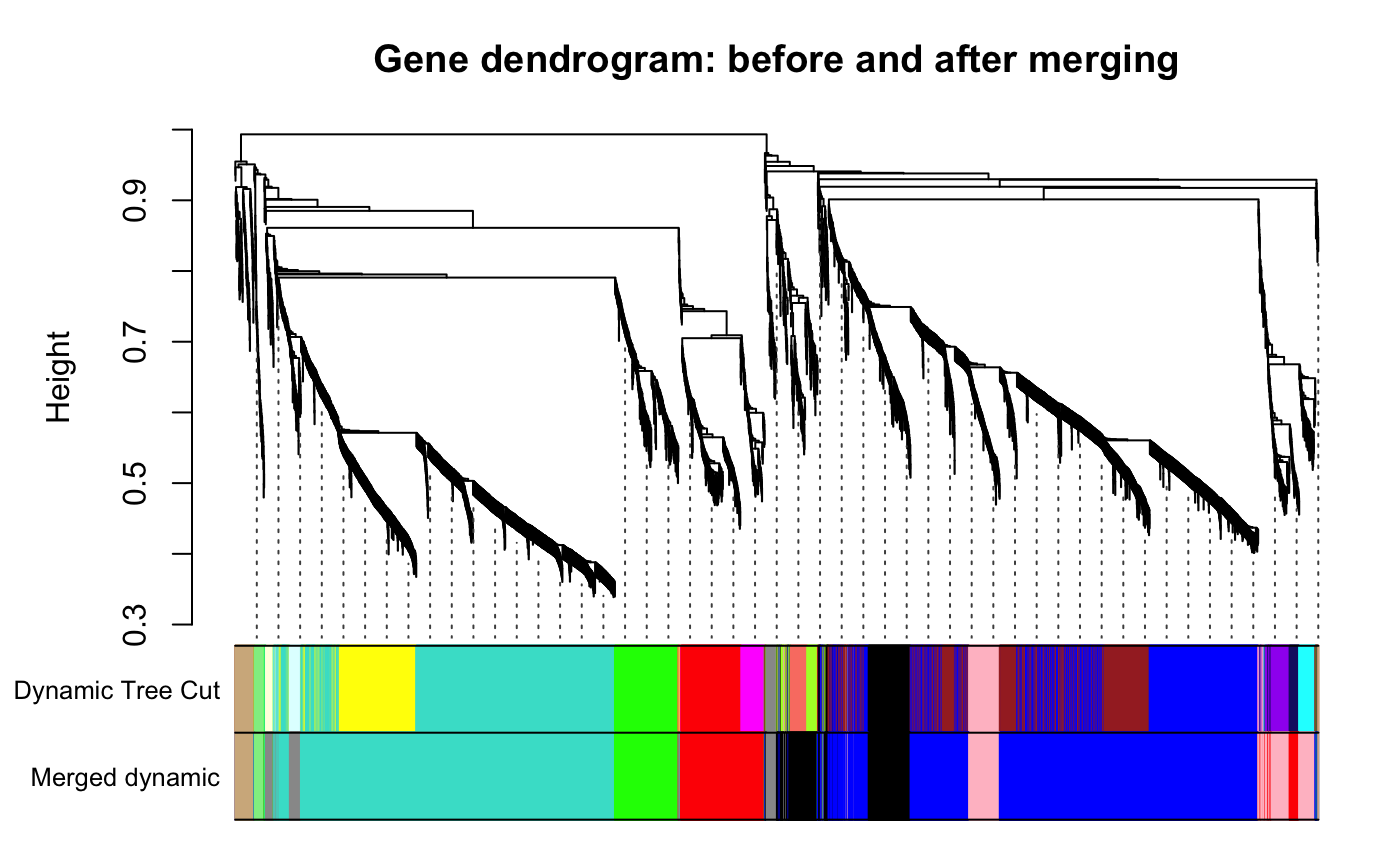
Applying Network-based clustering methods with WGCNA
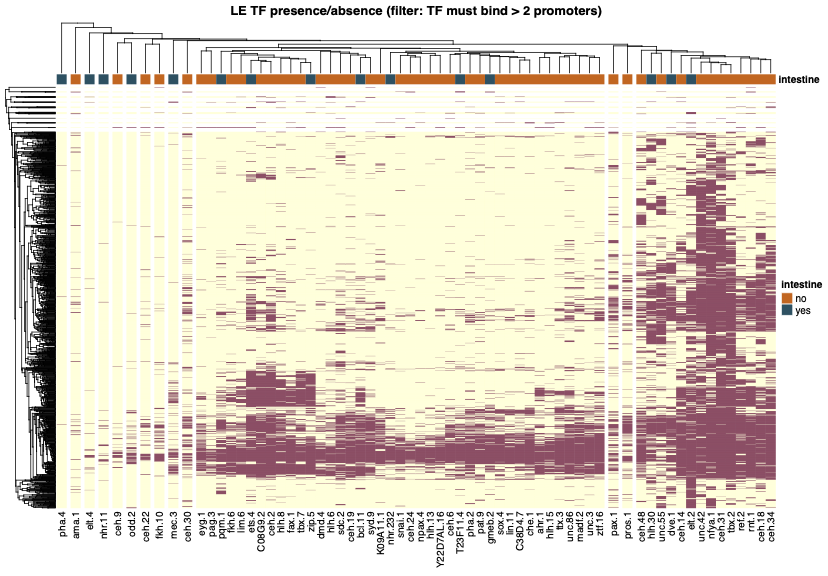
Clustering based on promoter binding profile of 96 Transcription Factors
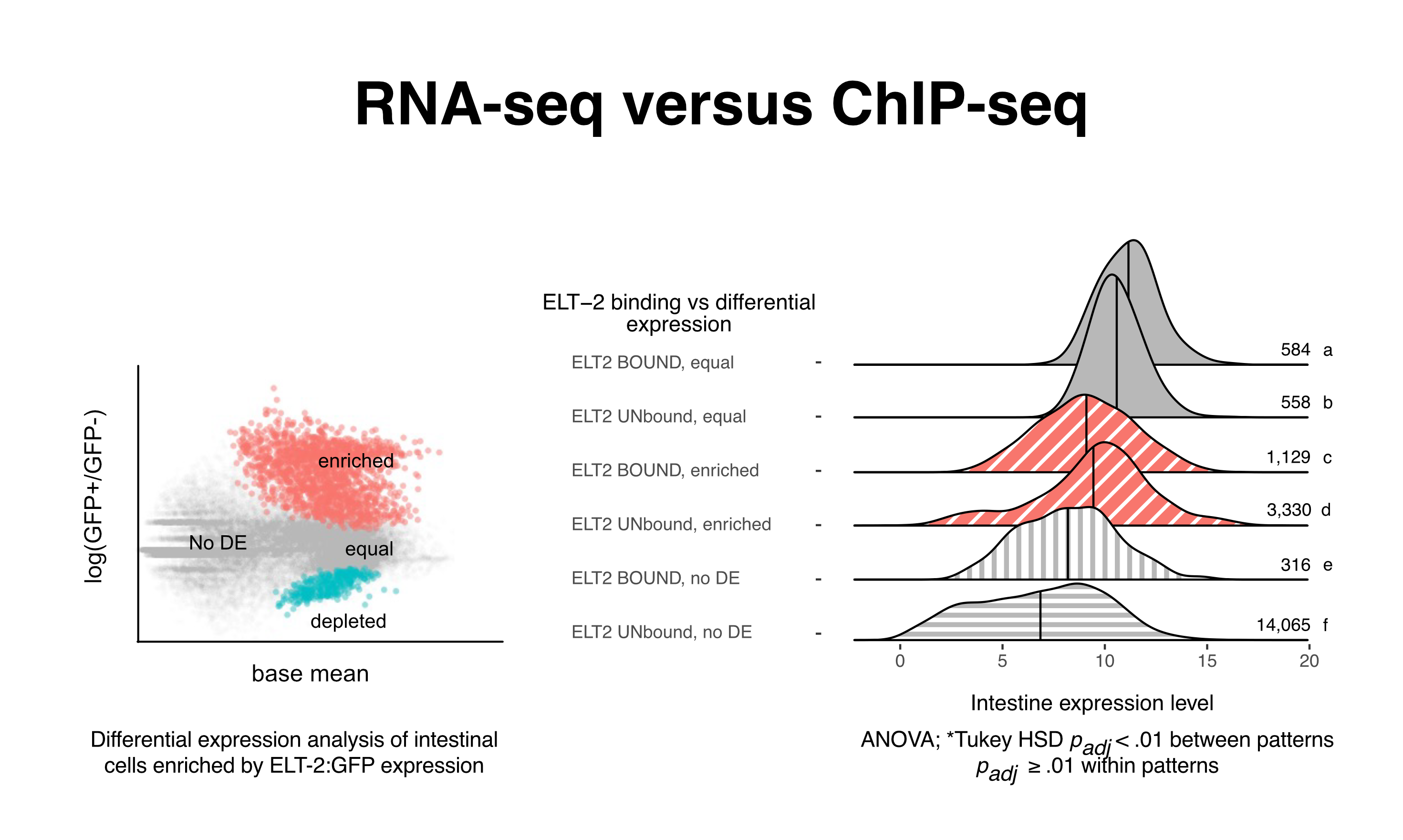
Comparison of occupancy score distributions for different gene expression classes
Recent Development
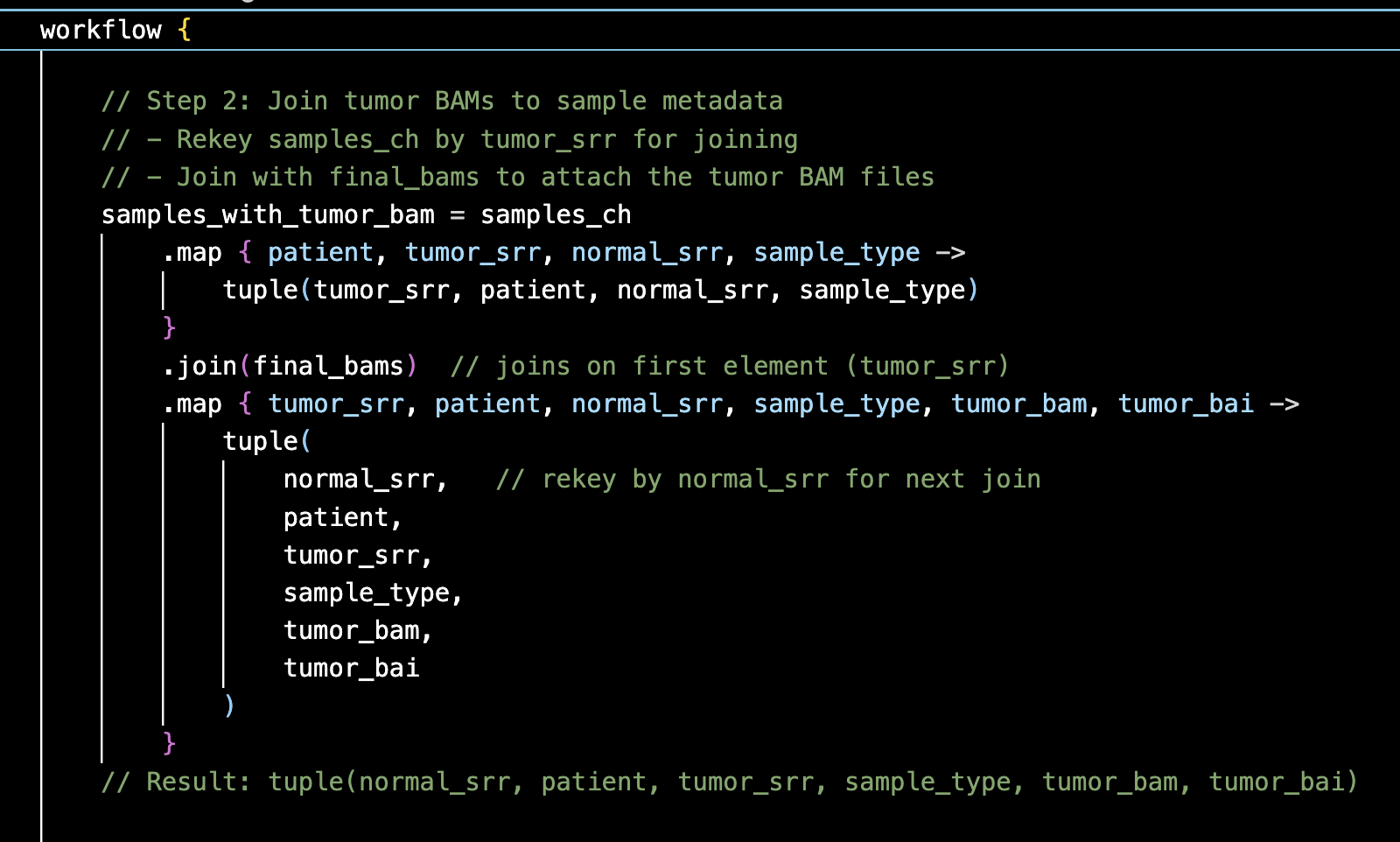
Tumor-normal somatic variant calling in colorectal cancer — pipeline tracks variants at targeted loci in patients as tumors become treatment-resistant across time points.
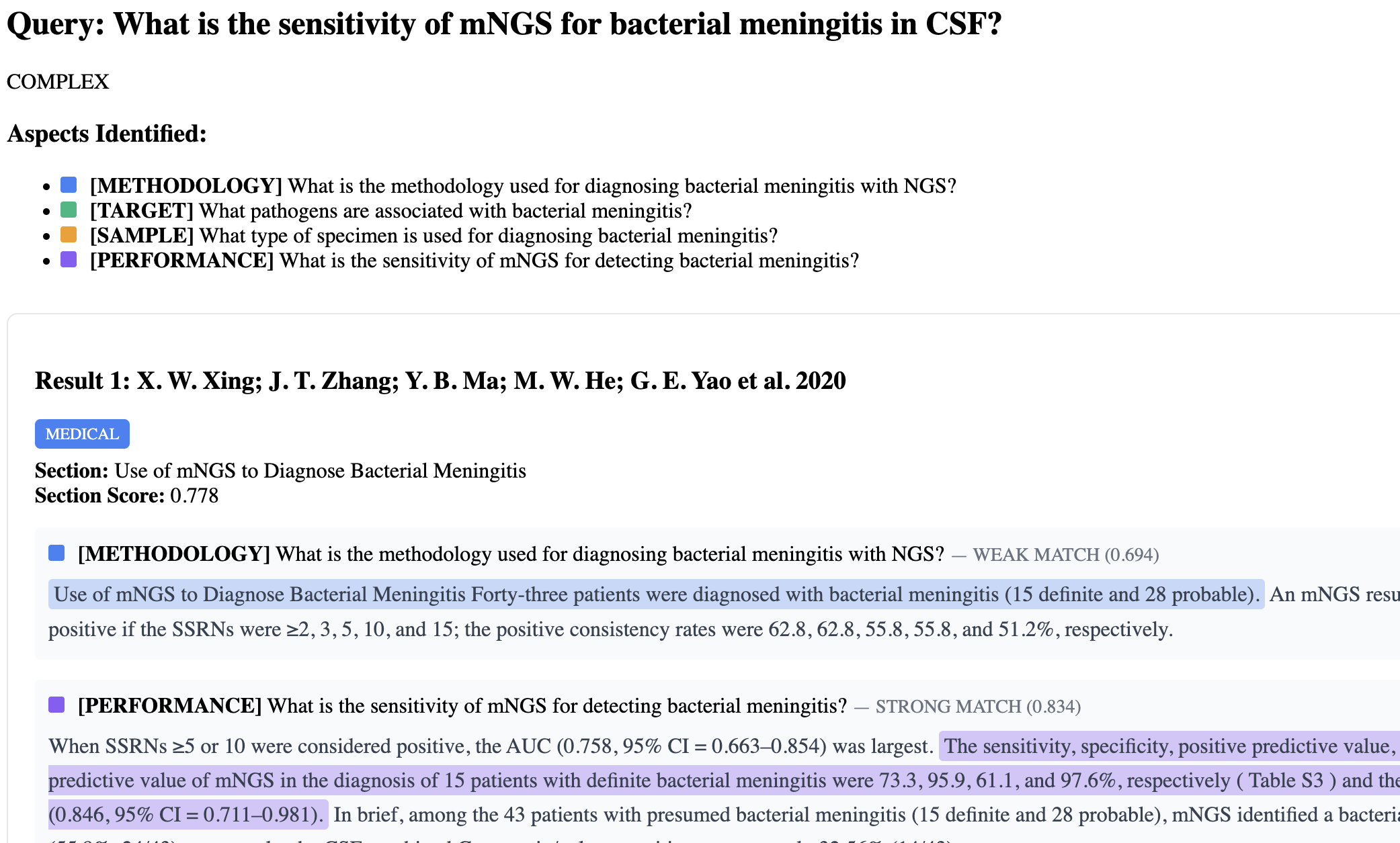
Custom AI search tool for NGS diagnostic literature — query the evidence base for sensitivity, specificity, and methodology across published studies.
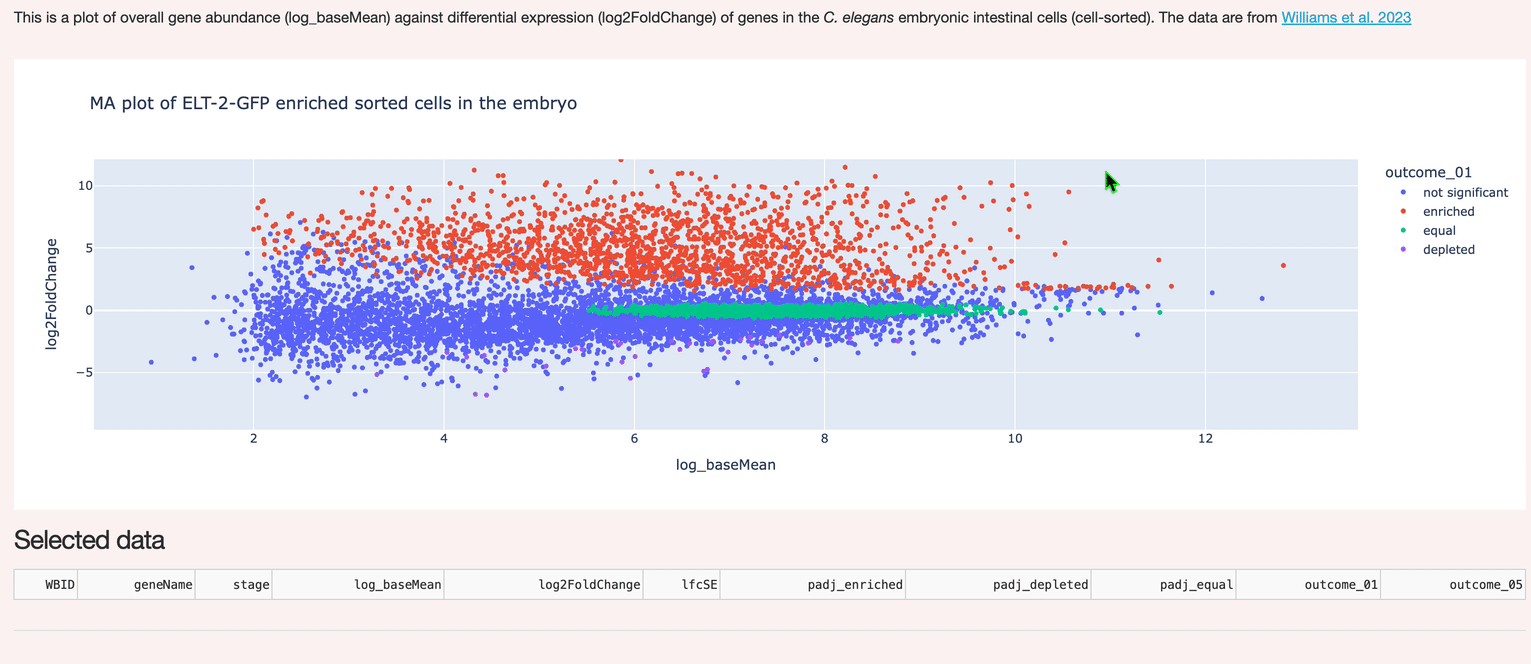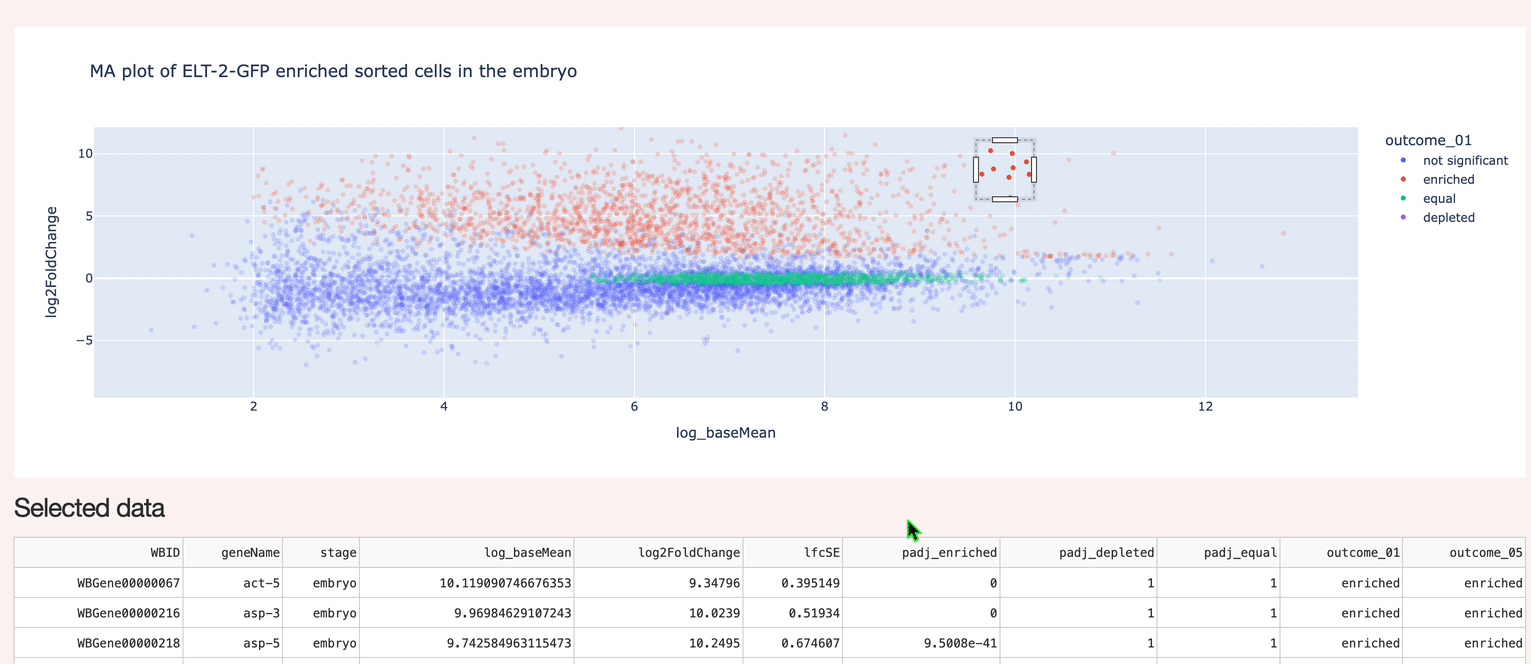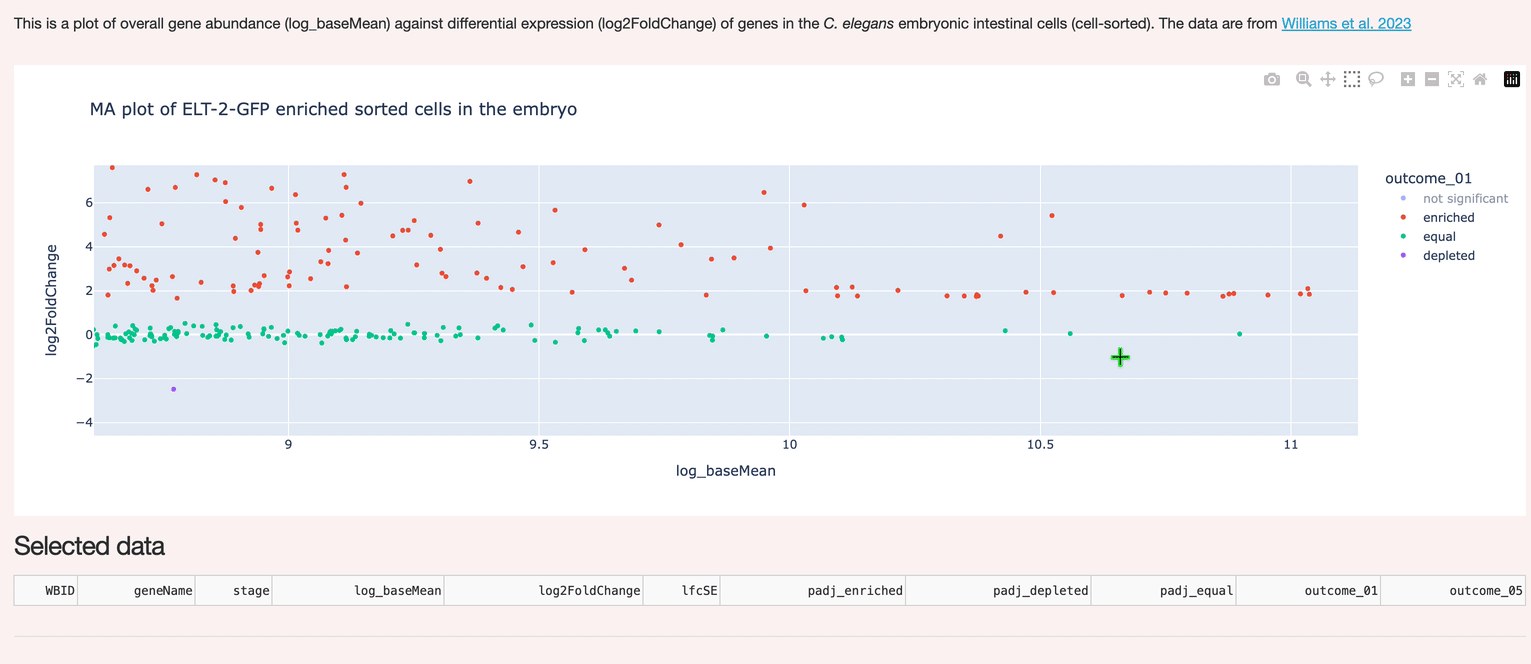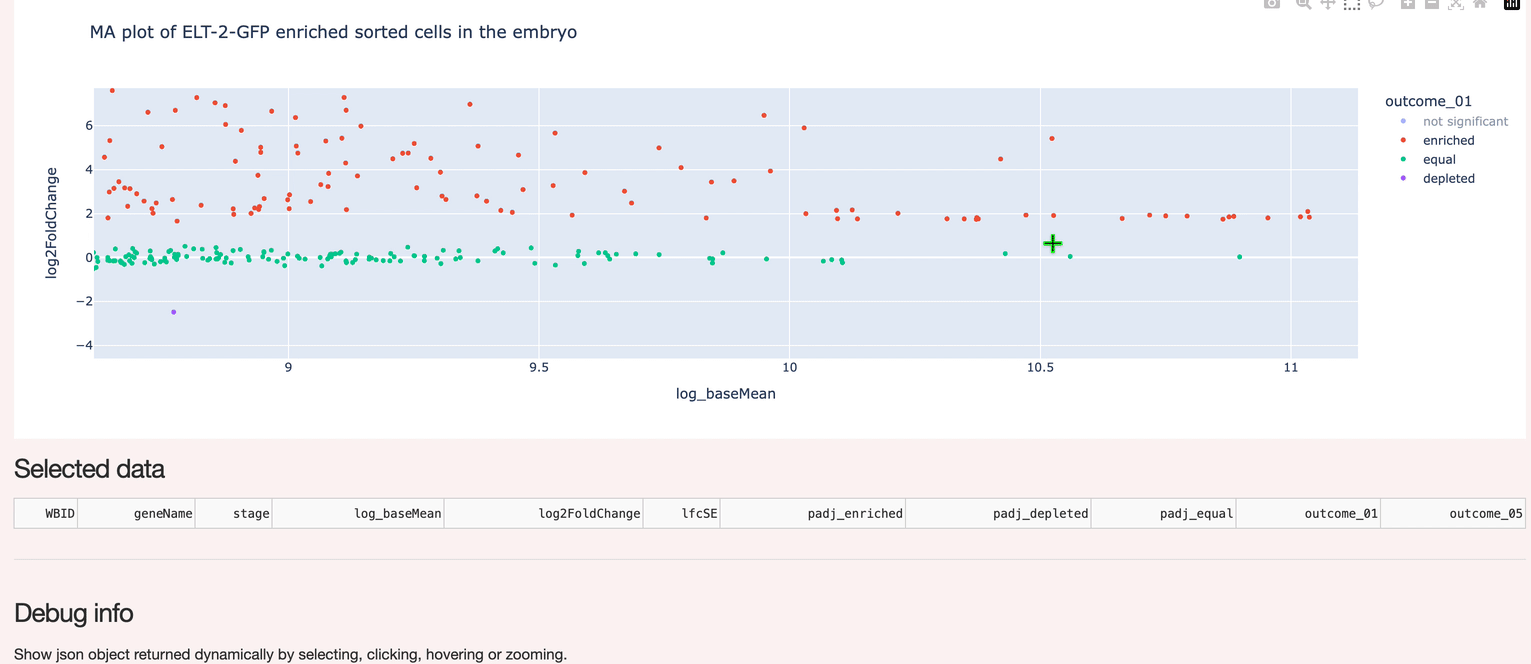
A custom interactive interface to hosted data (Postgres). Supports selection and inspection of gene expression data.
Cloud computing and training
Interactive learning environments for bioinformatics and computational biology